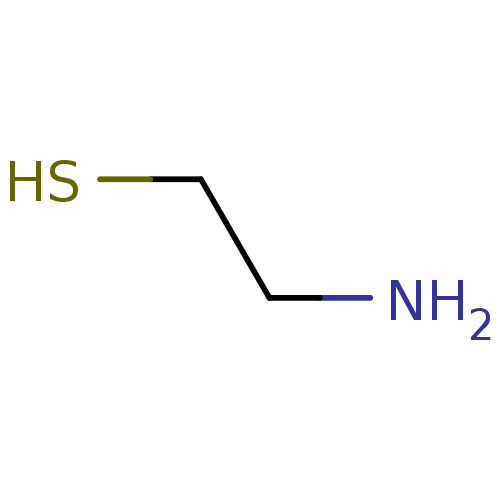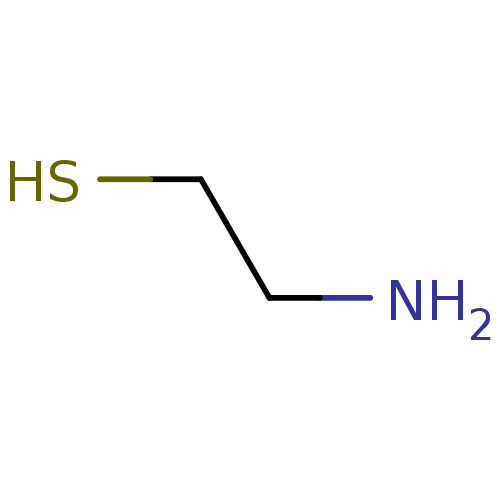 Cysteamine 2-aminoethane-1-thiol BDBM7968
Cysteamine 2-aminoethane-1-thiol BDBM7968
- Radwan, MO; Ciftci, HI; Ali, TFS; Koga, R; Tateishi, H; Nakata, A; Ito, A; Yoshida, M; Fujita, M; Otsuka, M Structure activity study of S-trityl-cysteamine dimethylaminopyridine derivatives as SIRT2 inhibitors: Improvement of SIRT2 binding and inhibition. Bioorg Med Chem Lett 30: (2020)
- Inhbition Assay The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 ug/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature.Five uL test compound (final concentration 0.1 nM to 100 uM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 uL of DPPI in MES buffer (final concentration 0.0125 ng/uL) and incubated for 10 min. Then, 5 uL of substrate in MES buffer (final concentration 50 uM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 uL of Gly-Phe-DMK in MES-buffer (final concentration 1 uM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm).
- Inhibition Assay The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 ug/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature.Five uL test compound (final concentration 0.1 nM to 100 uM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 uL of DPPI in MES buffer (final concentration 0.0125 ng/uL) and incubated for 10 min. Then, 5 uL of substrate in MES buffer (final concentration 50 uM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 uL of Gly-Phe-DMK in MES-buffer (final concentration 1 uM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm).
- Inhibition Assay The following buffers were used: MES buffer: 25 mM MES, 50 mM NaCl, 5 mM DTT, adjusted to pH 6.0, containing 0.1% BSA; TAGZyme Buffer: 20 mM NaH2P04, 150 mM NaCl adjusted to pH 6.0 with HC1Assay conditions: The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 ug/ml, respectively), and then activated by mixing in a 1 :2 ratio with a Cysteamine aqueous solution ( 2mM) and incubating for 5 min at room temperature.Five uL test compound (final concentration 0.1 nM to 100 uM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 uM of DPPI in MES buffer (final concentration 0.0125 ng/uL) and incubated for 10 min. Then, 5 uL of substrate in MES buffer (final concentration 50 uM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 uL of Gly-Phe-DMK in MES-buffer (final concentration 1 uM).
- Inhibition Assay The following buffers were used: MES buffer: 25 mM MES, 50 mM NaCl, 5 mM DTT, adjusted to pH 6.0, containing 0.1% BSA; TAGZyme Buffer: 20 mM NaH2PO4, 150 mM NaCl adjusted to pH 6.0 with HCl. Assay Conditions: The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 ug/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature.Five uL test compound (final concentration 0.1 nM to 100 uM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 uL of DPPI in MES buffer (final concentration 0.0125 ng/uL) and incubated for 10 min. Then, 5 uL of substrate in MES buffer (final concentration 50 uM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 uL of Gly-Phe-DMK in MES-buffer (final concentration 1 uM).
- Inhibition Assay The following buffers were used: MES buffer: 25 mM MES, 50 mM NaCl, 5 mM DTT, adjusted to pH 6.0, containing 0.1% BSA; TAGZyme Buffer: 20 mM NaH2PO4, 150 mM NaCl adjusted to pH is 6.0 with HClAssay Conditions:The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 ug/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 mM at room temperature.Five uL test compound (final concentration 0.1 nM to 100 uM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 uL of DPPI in MES buffer (final concentration 0.0125 ng/uL) and incubated for 10 min. Then, 5 uL of substrate in MES buffer (final concentration 50 uM) were added. The microtiter plates were then incubated at room temperature for 30 mM Then, the reaction was stopped by adding 10 uL of Gly-Phe-DMK in MES-buffer (final concentration 1 uM).
- Enzyme Assay The test inhibitors were solubilized in DMSO to a stock concentration of 30 mM. On the day of the assay, dose response plates were prepared by diluting the inhibitors in DMSO at compound concentration 200-fold the final in-assay concentration. Intermediate concentrations were prepared by diluting in DMSO in a four-fold series for a total of 11 data points.To prepare a working solution of human vanin-1, the enzyme was diluted to 33.3 pM in the assay buffer consisting of 50 mM Tris-HCl pH=8.0, 50 mM KCl, 0.005% Brij-35 and 1.6 mM cysteamine. To begin the assay 100 nL was transferred from the compound plate to the assay plate. Next, 15 μL of the vanin-1 working solution were transferred to the assay plate. The inhibitor and enzyme were incubated at room temperature for 30 minutes. The enzyme reaction was then initiated by the addition of 5 μL of 200 μM pantetheine 7-amino-4-trifluoromethylcoumarin prepared in assay buffer. The final concentrations in the assay were 25 pM human vanin-1 and 50 uM substrate. The final concentration of DMSO was 0.5%. The assay plates were incubated for 60 minutes and before they were read on a Perkin Elmer EnVision Model 2103 using a 405 nm excitation wavelength and a 510 nm emission wavelength for detection.
- Inhibition of Human DPPI (Cathepsin C) The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 μg/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature.Five uL test compound (final concentration 0.1 nM to 100 μM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 μL of DPPI in MES buffer (final concentration 0.0125 ng/L) and incubated for 10 min. Then, 5 μL of substrate in MES buffer (final concentration 50 μM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 μL of Gly-Phe-DMK in MES-buffer (final concentration 1 μM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm).Each assay microtiter plate contained wells with vehicle controls (1% DMSO in bidest+0.075% BSA) as reference for non-inhibited enzyme activity (100% Ctl; high values) and wells with inhibitor (Gly-Phe-DMK, in bidest+1% DMSO+0.075% BSA, final concentration 1 μM) as controls for background fluorescence (0% Ctl; low values).
- Inhibition of Human DPPI (Cathepsin C) The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 μg/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature. Five uL test compound (final concentration 0.1 nM to 100 μM) in aqua bidest 5 (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 μL of DPPI in MES buffer (final concentration 0.0125 ng/μL) and incubated for 10 min. Then, 5 μL of substrate in MES buffer (final concentration 50 μM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 μL of Gly-Phe-DMK in 10 MES-buffer (final concentration 1 μM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm).Each assay microtiter plate contained wells with vehicle controls (1% DMSO in bidest+0.075% 15 BSA) as reference for non-inhibited enzyme activity (100% Ctl; high values) and wells with inhibitor (Gly-Phe-DMK, in bidest+1% DMSO+0.075% BSA, final concentration 1 μM) as controls for background fluorescence (0% Ctl; low values). The analysis of the data was performed by calculating the percentage of fluorescence in the presence of test compound in comparison to the fluorescence of the vehicle control after 20 subtracting the background fluorescence using the following formula:(RFU(sample)−RFU(background))*100/(RFU(control)−RFU(background))
- Inhibition of Human DPPI Assay conditions: The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 μg/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2mM) and incubating for 5 min at room temperature.Five uL test compound (final concentration 0.1 nM to 100 μM) in aqua bidest 5 (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 μL of DPPI in MES buffer (final concentration 0.0125 ng/μL) and incubated for 10 min. Then, 5 μL of substrate in MES buffer (final concentration 50 μM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 μL of Gly-Phe-DMK in 10 MES-buffer (final concentration 1 μM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm).Each assay microtiter plate contained wells with vehicle controls (1% DMSO in bidest+0.075% 15 BSA) as reference for non-inhibited enzyme activity (100% Ctl; high values) and wells with inhibitor (Gly-Phe-DMK, in bidest+1% DMSO+0.075% BSA, final concentration 1 μM) as controls for background fluorescence (0% Ctl; low values). The analysis of the data was performed by calculating the percentage of fluorescence in the presence of test compound in comparison to the fluorescence of the vehicle control after 20 subtracting the background fluorescence using the following formula:(RFU(sample)−RFU(background))*100/(RFU(control)−RFU(background)
- Human Vanin-1 Enzyme Assay 1 The vanin-1 protein was prepared in-house from a construct expressing the extracellular domain of human vanin-1 (GenBank ID NM_004666) preceded N-terminally by the honey bee melittin signal peptide, a GSG linker sequence, a His6X tag and a FLAG tag. The secreted, soluble enzyme was purified from the conditioned medium from a CHO cell line stably expressing the resulting protein. Enzyme purification was performed through sequential Ni NTA and size-exclusion chromatography steps.The test inhibitors were solubilized in DMSO to a stock concentration of 30 mM. On the day of the assay, dose response plates were prepared by diluting the inhibitors in DMSO at compound concentration 200-fold the final in-assay concentration. Intermediate concentrations were prepared by diluting in DMSO in a four-fold series for a total of 11 data points.To prepare a working solution of human vanin-1, the enzyme was diluted to 33.3 pM in the assay buffer consisting of 50 mM Tris-HCl pH=8.0, 50 mM KCl, 0.005% Brij-35 and 1.6 mM cysteamine. To begin the assay 100 nL was transferred from the compound plate to the assay plate. Next, 15 μL of the vanin-1 working solution were transferred to the assay plate. The inhibitor and enzyme were incubated at room temperature for 30 minutes. The enzyme reaction was then initiated by the addition of 5 μL of 200 μM pantetheine 7-amino-4-trifluoromethylcoumarin prepared in assay buffer. The final concentrations in the assay were 25 pM human vanin-1 and 50 uM substrate. The final concentration of DMSO was 0.5%. The assay plates were incubated for 60 minutes and before they were read on a Perkin Elmer EnVision Model 2103 using a 405 nm excitation wavelength and a 510 nm emission wavelength for detection.
- Human Vanin-1 Enzyme Assay 2 The vanin-1 protein was prepared in-house from a construct expressing the extracellular domain of human vanin-1 (GenBank ID NM_004666) preceded N-terminally by the honey bee melittin signal peptide, a GSG linker sequence, a His6X tag and a FLAG tag. The secreted, soluble enzyme was purified from the conditioned medium from a CHO cell line stably expressing the resulting protein. Enzyme purification was performed through sequential Ni NTA and size-exclusion chromatography steps.On the day of the assay, dose response plates were prepared by diluting the inhibitors in DMSO at compound concentration 100-fold the final in-assay concentration. Concentration series were prepared by serially diluting in DMSO in a half-log series for a total of 11 data points. Intermediate compound plates containing compound in 10% DMSO were then created by diluting the compounds 10-fold in assay buffer consisting of 50 mM Tris-HCl pH=8.0, 50 mM KCl, 0.005% Brij-35 and 1.5 mM cysteamine. To begin the assay 3 μL were transferred from the intermediate compound plate to the assay plate.A working solution of human vanin-1 was prepared by diluting the enzyme stock to 1.25 nM in assay buffer. Next, 24 μL of the vanin-1 working solution were transferred to the assay plate. The enzyme reaction was then initiated by the addition of 3 μL of 100 μM pantetheine 7-amino-4-trifluoromethylcoumarin prepared in assay buffer. The final concentrations in the assay were 1 nM human vanin-1 and 10 uM substrate. The final concentration of DMSO was 1%. The assay plates were incubated for 45 minutes and before they were read on a Spectramax M5 using a 405 nm excitation wavelength and a 505 nm emission wavelength for detection.
- Human Vanin-1 Enzyme Assay 3 The vanin-1 protein was prepared in-house from a construct expressing the extracellular domain of human vanin-1 (GenBank ID NM_004666) preceded N-terminally by the honey bee melittin signal peptide, a GSG linker sequence, a His6X tag and a FLAG tag. The secreted, soluble enzyme was purified from the conditioned medium from a CHO cell line stably expressing the resulting protein. Enzyme purification was performed through sequential Ni NTA and size-exclusion chromatography steps.On the day of the assay, dose response plates were prepared by diluting the inhibitors in DMSO at compound concentration 100-fold the final in-assay concentration. Concentration series were prepared by serially diluting in DMSO in a two-fold series for a total of 11 data points. Intermediate compound plates containing compound in 10% DMSO were then created by diluting the compounds 10-fold in assay buffer consisting of 50 mM Tris-HCl pH=8.0, 50 mM KCl, 0.005% Brij-35 and 1.5 mM cysteamine. To begin the assay 3 μL were transferred from the intermediate compound plate to the assay plate.A working solution of human vanin-1 was prepared by diluting the enzyme stock to 2.5 nM in assay buffer. Next, 24 μL of the vanin-1 working solution were transferred to the assay plate. The enzyme reaction was then initiated by the addition of 3 μL of 100 μM pantetheine 7-amino-4-trifluoromethylcoumarin prepared in 5 uM acetic acid. The final concentrations in the assay were 2 nM human vanin-1 and 10 uM substrate. The final concentration of DMSO was 1%. The assay plates were incubated at room temperature for 15 minutes and before they were read on a Tecan Safire using a 405 nm excitation wavelength and a 505 nm emission wavelength for detection.
- Inhibition of Human DPPI (Cathepsin C) Materials: Microtiterplates (Optiplate-384 F) were purchased from PerkinElmer (Prod. No. 10 6007270). The substrate Gly-Arg-AMC was from Biotrend (Prod.-No. 808756 Custom peptide).Bovine serum albumin (BSA; Prod. No. A3059) and Dithiothreitol (DTT; Prod. No D0632) were from Sigma. TagZyme buffer was from Riedel-de-Haen (Prod.-No. 04269), NaCl was from Merck (Prod.-No. 1.06404.1000) and morpholinoethane sulfonic acid (MES), was from Serva (Prod.-No. 29834). The DPP1 inhibitor Gly-Phe-DMK was purchased from MP Biomedicals (Prod.-No. 03DK00625). The recombinant human DPPI was purchased from Prozymex. All other materials were of highest grade commercially available.The following buffers were used: MES buffer: 25 mM MES, 50 mM NaCl, 5 mM DTT, adjusted to pH 6.0, containing 0.1% BSA; TAGZyme Buffer: 20 mM NaH2PO4, 150 mM NaCl adjusted to pH 20 6.0 with HClAssay Conditions:The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 μg/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature. Five uL test compound (final concentration 0.1 nM to 100 μM) in aqua bidest 5 (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 μL of DPPI in MES buffer (final concentration 0.0125 ng/μL) and incubated for 10 min. Then, 5 μL of substrate in MES buffer (final concentration 50 μM) were added. The microtiter plates were then incubated at room temperature for 30 min. Then, the reaction was stopped by adding 10 μL of Gly-Phe-DMK in 10 MES-buffer (final concentration 1 μM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm).Each assay microtiter plate contained wells with vehicle controls (1% DMSO in bidest+0.075% 15 BSA) as reference for non-inhibited enzyme activity (100% Ctl; high values) and wells with inhibitor (Gly-Phe-DMK, in bidest+1% DMSO+0.075% BSA, final concentration 1 μM) as controls for background fluorescence (0% Ctl; low values).
- Inhibition Assay of Human DPPI (Cathepsin C) Materials: Microtiterplates (Optiplate-384 F) were purchased from PerkinElmer (Prod. No. 6007270). The substrate Gly-Arg-AMC was from Biotrend (Prod.-No. 808756 Custom peptide). Bovine serum albumin (BSA; Prod. No. A3059) and Dithiothreitol (DTT; Prod. No D0632) were from Sigma. TagZyme buffer was from Riedel-de-Haen (Prod.-No. 04269), NaCl was from Merck (Prod.-No. 1.06404.1000) and morpholinoethane sulfonic acid (MES), was from Serva (Prod.-No. 29834). The DPP1 inhibitor Gly-Phe-DMK was purchased from MP Biomedicals (Prod.-No. 03DK00625). The recombinant human DPPI was purchased from Prozymex. All other materials were of highest grade commercially available. The following buffers were used: MES buffer: 25 mM MES, 50 mM NaCl, 5 mM DTT, adjusted to pH 6.0, containing 0.1% BSA; TAGZyme Buffer: 20 mM NaH2PO4, 150 mM NaCl adjusted to pH 6.0 with HCl. Assay conditions: The recombinant human DPPI was diluted in TAGZyme buffer to 1 U/ml (38.1 μg/ml, respectively), and then activated by mixing in a 1:2 ratio with a Cysteamine aqueous solution (2 mM) and incubating for 5 min at room temperature. Five uL test compound (final concentration 0.1 nM to 100 μM) in aqua bidest (containing 4% DMSO, final DMSO concentration 1%) were mixed with 10 μL of DPPI in MES buffer (final concentration 0.0125 ng/μL) and incubated for 10 min. Then, 5 μL of substrate in MES buffer (final concentration 50 μM) were added. The microtiter plates were then incubated at room temperature for 30 mM Then, the reaction was stopped by adding 10 μL of Gly-Phe-DMK in MES-buffer (final concentration 1 μM). The fluorescence in the wells was determined using a Molecular Devices SpectraMax M5 Fluorescence Reader (Ex 360 nm, Em 460 nm) or an Envision Fluorescence Reader (Ex 355 nm, Em 460 nm). Each assay microtiter plate contained wells with vehicle controls (1% DMSO in bidest+0.075% BSA) as reference for non-inhibited enzyme activity (100% Ctl; high values) and wells with inhibitor (Gly-Phe-DMK, in bidest+1% DMSO+0.075% BSA, final concentration 1 μM) as controls for background fluorescence (0% Ctl; low values). The analysis of the data was performed by calculating the percentage of fluorescence in the presence of test compound in comparison to the fluorescence of the vehicle control after subtracting the background fluorescence using the following formula: (RFU(sample)−RFU(background))*100/(RFU(control)−RFU(background)).